
The trefoil knot fold is a protein fold in which the protein backbone is twisted into a trefoil knot shape. "Shallow" knots in which the tail of the polypeptide chain only passes through a loop by a few residues are uncommon, but "deep" knots in which many residues are passed through the loop are extremely rare. Deep trefoil knots have been found in the SPOUT superfamily.[1] including methyltransferase proteins involved in posttranscriptional RNA modification in all three domains of life, including bacterium Thermus thermophilus[2] and proteins,[3] in archaea[1] and in eukaryota.[4]
In many cases the trefoil knot is part of the active site or a ligand-binding site and is critical to the activity of the enzyme in which it appears. Before the discovery of the first knotted protein, it was believed that the process of protein folding could not efficiently produce deep knots in protein backbones. Studies of the folding kinetics of a dimeric protein from Haemophilus influenzae have revealed that the folding of trefoil knot proteins may depend on proline isomerization.[5] Computational algorithms have been developed to identify knotted protein structures, both to canvas the Protein Data Bank for previously undetected natural knots and to identify knots in protein structure predictions, where they are unlikely to accurately reproduce the native-state structure due to the rarity of knots in known proteins.[6] Knottins are small, diverse and stable proteins with important drug design potential. They can be classified in 30 families which cover a wide range of sequences (1621 sequenced), three-dimensional structures (155 solved) and functions (> 10). Inter knottin similarity lies mainly between 20% and 40% sequence identity and 1.5 to 4 A backbone deviations although they all share a tightly knotted disulfide core. This important variability is likely to arise from the highly diverse loops which connect the successive knotted cysteines. The prediction of structural models for all knottin sequences would open new directions for the analysis of interaction sites and to provide a better understanding of the structural and functional organization of proteins sharing this scaffold.[7]
Trefoil domain
| Trefoil (P-type) domain | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
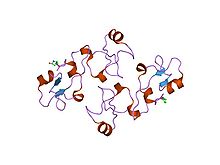 Structure of pancreatic spasmolytic polypeptide.[8] | |||||||||||
| Identifiers | |||||||||||
| Symbol | Trefoil | ||||||||||
| Pfam | PF00088 | ||||||||||
| InterPro | IPR000519 | ||||||||||
| SMART | SM00018 | ||||||||||
| PROSITE | PDOC00024 | ||||||||||
| SCOP2 | 1psp / SCOPe / SUPFAM | ||||||||||
| CDD | cd00111 | ||||||||||
| |||||||||||
Trefoil (P-type) domain is a cysteine-rich domain of approximately forty five amino-acid residues has been found in some extracellular eukaryotic proteins.[9][10][11][12] It is known as either the 'P', 'trefoil' or 'TFF' domain, and contains six cysteines linked by three disulphide bonds with connectivity 1–5, 2–4, 3–6.
The domain has been found in a variety of extracellular eukaryotic proteins,[9][11][12] including protein pS2 (TFF1) a protein secreted by the stomach mucosa; spasmolytic polypeptide (SP) (TFF2), a protein of about 115 residues that inhibits gastrointestinal motility and gastric acid secretion; intestinal trefoil factor (ITF) (TFF3); Xenopus laevis stomach proteins xP1 and xP4; xenopus integumentary mucins A.1 (preprospasmolysin) and C.1, proteins which may be involved in defense against microbial infections by protecting the epithelia from the external environment; xenopus skin protein xp2 (or APEG); Zona pellucida sperm-binding protein B (ZP-B); intestinal sucrase-isomaltase (EC 3.2.1.48 / EC 3.2.1.10), a vertebrate membrane bound, multifunctional enzyme complex which hydrolyzes sucrose, maltose and isomaltose; and lysosomal alpha-glucosidase (EC 3.2.1.20).
Examples
Human gene encoding proteins containing the trefoil domain include:
History
There was a web server pKNOT available to detect knots in proteins as well as to provide information on knotted proteins in the Protein Data Bank.[13]
References
- ^ Zarembinski, Thomas I.; Kim, Youngchang; Peterson, Kelly; Christendat, Dinesh; Dharamsi, Akil; Arrowsmith, Cheryl H.; Edwards, Aled M.; Joachimiak, Andrzej (2002). "Deep trefoil knot implicated in RNA binding found in an archaebacterial protein". Proteins: Structure, Function, and Bioinformatics. 50 (2): 177–183. doi:10.1002/prot.10311. PMC 2792022. PMID 12486711.
- ^ Nureki, Osamu; Shirouzu, Mikako; Hashimoto, Kyoko; Ishitani, Ryuichiro; Terada, Takaho; Tamakoshi, Masatada; Oshima, Tairo; Chijimatsu, Masao; Takio, Koji; Vassylyev, Dmitry G.; Shibata, Takehiko; Inoue, Yorinao; Kuramitsu, Seiki; Yokoyama, Shigeyuki (2002). "An enzyme with a deep trefoil knot for the active-site architecture". Acta Crystallographica Section D Biological Crystallography. 58 (7): 1129–1137. doi:10.1107/s0907444902006601. PMID 12077432.
- ^ Nureki, Osamu; Watanabe, Kazunori; Fukai, Shuya; Ishii, Ryohei; Endo, Yaeta; Hori, Hiroyuki; Yokoyama, Shigeyuki (2004). "Deep Knot Structure for Construction of Active Site and Cofactor Binding Site of tRNA Modification Enzyme". Structure. 12 (4): 593–602. doi:10.1016/j.str.2004.03.003. PMID 15062082.
- ^ Leulliot, N.; Bohnsack, M. T.; Graille, M.; Tollervey, D.; Van Tilbeurgh, H. (2007). "The yeast ribosome synthesis factor Emg1 is a novel member of the superfamily of alpha/Beta knot fold methyltransferases". Nucleic Acids Research. 36 (2): 629–639. doi:10.1093/nar/gkm1074. PMC 2241868. PMID 18063569.
- ^ Mallam, Anna L.; Jackson, Sophie E. (2006). "Probing Nature's Knots: The Folding Pathway of a Knotted Homodimeric Protein". Journal of Molecular Biology. 359 (5): 1420–1436. doi:10.1016/j.jmb.2006.04.032. PMID 16787779.
- ^ Khatib, Firas; Weirauch, Matthew T.; Rohl, Carol A. (2006). "Rapid knot detection and application to protein structure prediction". Bioinformatics. 22 (14): e252–e259. doi:10.1093/bioinformatics/btl236. PMID 16873480.
- ^ Gracy, Jérôme; Chiche, Laurent (2010). "Optimizing structural modeling for a specific protein scaffold: Knottins or inhibitor cystine knots". BMC Bioinformatics. 11: 535. doi:10.1186/1471-2105-11-535. PMC 2984590. PMID 21029427.
- ^ Gajhede M, Petersen TN, Henriksen A, et al. (December 1993). "Pancreatic spasmolytic polypeptide: first three-dimensional structure of a member of the mammalian trefoil family of peptides". Structure. 1 (4): 253–62. doi:10.1016/0969-2126(93)90014-8. PMID 8081739.
- ^ a b Otto B, Wright N (1994). "Trefoil peptides. Coming up clover". Current Biology. 4 (9): 835–838. doi:10.1016/S0960-9822(00)00186-X. PMID 7820556. S2CID 11245174.
- ^ Thim L, Wright NA, Hoffmann W, Otto WR, Rio MC (1997). "Rolling in the clover: trefoil factor family (TFF)-domain peptides, cell migration and cancer". FEBS Letters. 408 (2): 121–123. doi:10.1016/S0014-5793(97)00424-9. PMID 9187350. S2CID 26946754.
- ^ a b Bork P (1993). "A trefoil domain in the major rabbit zona pellucida protein". Protein Science. 2 (4): 669–670. doi:10.1002/pro.5560020417. PMC 2142363. PMID 8518738.
- ^ a b Hoffmann W, Hauser F (1993). "The P-domain or trefoil motif: a role in renewal and pathology of mucous epithelia?". Trends in Biochemical Sciences. 18 (7): 239–243. doi:10.1016/0968-0004(93)90170-R. PMID 8267796.
- ^ Lai, Y.-L.; Yen, S.-C.; Yu, S.-H.; Hwang, J.-K. (2007). "pKNOT: The protein KNOT web server". Nucleic Acids Research. 35 (Web Server issue): W420–W424. doi:10.1093/nar/gkm304. PMC 1933195. PMID 17526524.
External links
Bibliography
- Tkaczuk KL, Dunin-Horkawicz S, Purta E, Bujnicki JM. (2007). Structural and evolutionary bioinformatics of the SPOUT superfamily of methyltransferases. BMC Bioinformatics. 8:73








