Serine/threonine-protein kinase PLK4 also known as polo-like kinase 4 is an enzyme that in humans is encoded by the PLK4 gene.[5] The Drosophila homolog is SAK, the C. elegans homolog is zyg-1, and the Xenopus homolog is Plx4.[6]
Function
PLK4 encodes a member of the polo family of serine/threonine protein kinases. The protein localizes to centrioles—complex microtubule-based structures found in centrosomes—and regulates centriole duplication during the cell cycle.[5] Overexpression of PLK4 results in centrosome amplification, and knockdown of PLK4 results in loss of centrosomes.[7][8]
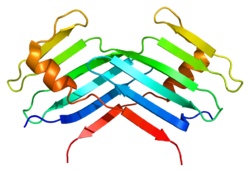
Structure
PLK4 contains an N-terminal kinase domain (residues 12-284) and a C-terminal localization domain (residues 596-898).[9] Other polo-like kinase members contain 2 C-terminal polo box domains (PBD). PLK4 contains these 2 domains in addition to a third PBD, which facilitates oligomerization, targeting, and promotes trans-autophosphorylation, limiting centriole duplication to once per cell cycle.[9]
As a cancer drug target
Inhibitors of the enzymatic activity PLK4 have potential in the treatment of cancer.[10][11] The PLK4 inhibitor R1530 down regulates the expression of mitotic checkpoint kinase BubR1 that in turn leads to polyploidy rendering cancer cells unstable and more sensitive to cancer chemotherapy. Furthermore, normal cells are resistant to the polyploidy inducing effects of R1530.[12]
Another PLK4 inhibitor, CFI-400945 has demonstrated efficacy in animal models of breast and ovarian cancer.[13][14]
Another PLK4 inhibitor, centrinone, has been reported to deplete centrioles in human and other vertebrate cell types, which resulted in a p53-dependent cell cycle arrest in G1.[15] Inhibition of PLK4 using a chemical genetic strategy has validated this p53-dependent cell cycle arrest in G1.[16]
PLK4 was also identified as a potential therapeutic target for malignant rhabdoid tumors, medulloblastomas and possibly, other embryonal tumors of the brain.[17][18][19][20]
Interactions and substrates
Documented PLK4 substrates include STIL, GCP6,[21] Hand1,[22][23] Ect2,[24] FBXW5,[25] and itself (via autophosphorylation). Autophosphorylation of PLK4 results in ubiquitination and subsequent destruction by the proteasome.[26][27]
PLK4 has been shown to interact with Stratifin.[28]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000142731 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000025758 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b "Entrez Gene: PLK4 polo-like kinase 4 (Drosophila)".
- ^ Shimanovskaya E, Viscardi V, Lesigang J, Lettman MM, Qiao R, Svergun DI, Round A, Oegema K, Dong G (August 2014). "Structure of the C. elegans ZYG-1 cryptic polo box suggests a conserved mechanism for centriolar docking of Plk4 kinases". Structure. 22 (8): 1090–1104. doi:10.1016/j.str.2014.05.009. PMC 4126857. PMID 24980795.
- ^ Godinho SA, Picone R, Burute M, Dagher R, Su Y, Leung CT, Polyak K, Brugge JS, Théry M, Pellman D (June 2014). "Oncogene-like induction of cellular invasion from centrosome amplification". Nature. 510 (7503): 167–71. Bibcode:2014Natur.510..167G. doi:10.1038/nature13277. PMC 4061398. PMID 24739973.
- ^ Habedanck R, Stierhof YD, Wilkinson CJ, Nigg EA (November 2005). "The Polo kinase Plk4 functions in centriole duplication". Nature Cell Biology. 7 (11): 1140–6. doi:10.1038/ncb1320. PMID 16244668. S2CID 1349505.
- ^ a b Slevin LK, Nye J, Pinkerton DC, Buster DW, Rogers GC, Slep KC (November 2012). "The structure of the plk4 cryptic polo box reveals two tandem polo boxes required for centriole duplication". Structure. 20 (11): 1905–17. doi:10.1016/j.str.2012.08.025. PMC 3496063. PMID 23000383.
- ^ Suri A, Bailey AW, Tavares MT, Gunosewoyo H, Dyer CP, Grupenmacher AT, Piper DR, Horton RA, Tomita T, Kozikowski AP, Roy SM, Sredni ST (April 2019). "Evaluation of Protein Kinase Inhibitors with PLK4 Cross-Over Potential in a Pre-Clinical Model of Cancer". International Journal of Molecular Sciences. 20 (9): 2112. doi:10.3390/ijms20092112. PMC 6540285. PMID 31035676.
- ^ Mason J, Wei S, Luo X, Nadeem V, Kiarash R, Huang P, Awrey D, Leung G, Beletskaya I, Feher M, Forrest B, Laufer R, Sampson P, Li SW, Liu Y, Lang Y, Pauls H, Mak T, Pan JG. "Inhibition of Polo-like kinase 4 as an anti-cancer strategy". Abstract LB-215. Cancer Research. pp. LB-215.
{{cite web}}: CS1 maint: deprecated archival service (link) - ^ Tovar C, Higgins B, Deo D, Kolinsky K, Liu JJ, Heimbrook DC, Vassilev LT (August 2010). "Small-molecule inducer of cancer cell polyploidy promotes apoptosis or senescence: Implications for therapy". Cell Cycle. 9 (16): 3364–75. doi:10.4161/cc.9.16.12732. PMID 20814247.
- ^ "Experimental drug shows promise in treating breast, ovarian cancer". News. Canadian Broadcasting Corporation. 2013-06-18.
- ^ Yu B, Yu Z, Qi PP, Yu DQ, Liu HM (May 2015). "Discovery of orally active anticancer candidate CFI-400945 derived from biologically promising spirooxindoles: success and challenges". European Journal of Medicinal Chemistry. 95: 35–40. doi:10.1016/j.ejmech.2015.03.020. PMID 25791677.
- ^ Wong YL, Anzola JV, Davis RL, Yoon M, Motamedi A, Kroll A, Seo CP, Hsia JE, Kim SK, Mitchell JW, Mitchell BJ, Desai A, Gahman TC, Shiau AK, Oegema K (June 2015). "Cell biology. Reversible centriole depletion with an inhibitor of Polo-like kinase 4". Science. 348 (6239): 1155–60. doi:10.1126/science.aaa5111. PMC 4764081. PMID 25931445.
- ^ Lambrus BG, Uetake Y, Clutario KM, Daggubati V, Snyder M, Sluder G, Holland AJ (July 2015). "p53 protects against genome instability following centriole duplication failure". The Journal of Cell Biology. 210 (1): 63–77. doi:10.1083/jcb.201502089. PMC 4494000. PMID 26150389.
- ^ Sredni ST, Suzuki M, Yang JP, Topczewski J, Bailey AW, Gokirmak T, Gross JN, de Andrade A, Kondo A, Piper DR, Tomita T (November 2017). "A functional screening of the kinome identifies the Polo-like kinase 4 as a potential therapeutic target for malignant rhabdoid tumors, and possibly, other embryonal tumors of the brain". Pediatric Blood & Cancer. 64 (11) e26551. doi:10.1002/pbc.26551. PMID 28398638.
- ^ Sredni ST, Bailey AW, Suri A, Hashizume R, He X, Louis N, Gokirmak T, Piper DR, Watterson DM, Tomita T (December 2017). "Inhibition of polo-like kinase 4 (PLK4): a new therapeutic option for rhabdoid tumors and pediatric medulloblastoma". Oncotarget. 8 (67): 111190–111212. doi:10.18632/oncotarget.22704. PMC 5762315. PMID 29340047.
- ^ Bailey AW, Suri A, Chou PM, Pundy T, Gadd S, Raimondi SL, Tomita T, Sredni ST (November 2018). "Polo-Like Kinase 4 (PLK4) Is Overexpressed in Central Nervous System Neuroblastoma (CNS-NB)". Bioengineering. 5 (4): 96. doi:10.3390/bioengineering5040096. PMC 6315664. PMID 30400339.
- ^ Suri A, Bailey AW, Tavares MT, Gunosewoyo H, Dyer CP, Grupenmacher AT, Piper DR, Horton RA, Tomita T, Kozikowski AP, Roy SM, Sredni ST (April 2019). "Evaluation of Protein Kinase Inhibitors with PLK4 Cross-Over Potential in a Pre-Clinical Model of Cancer". International Journal of Molecular Sciences. 20 (9): 2112. doi:10.3390/ijms20092112. PMC 6540285. PMID 31035676.
- ^ Bahtz R, Seidler J, Arnold M, Haselmann-Weiss U, Antony C, Lehmann WD, Hoffmann I (January 2012). "GCP6 is a substrate of Plk4 and required for centriole duplication". Journal of Cell Science. 125 (Pt 2): 486–96. doi:10.1242/jcs.093930. PMID 22302995.
- ^ Martindill DM, Risebro CA, Smart N, Franco-Viseras Mdel M, Rosario CO, Swallow CJ, Dennis JW, Riley PR (October 2007). "Nucleolar release of Hand1 acts as a molecular switch to determine cell fate". Nature Cell Biology. 9 (10): 1131–41. doi:10.1038/ncb1633. PMID 17891141. S2CID 5110337.
- ^ Hudson JW, Kozarova A, Cheung P, Macmillan JC, Swallow CJ, Cross JC, Dennis JW (March 2001). "Late mitotic failure in mice lacking Sak, a polo-like kinase". Current Biology. 11 (6): 441–6. Bibcode:2001CBio...11..441H. doi:10.1016/s0960-9822(01)00117-8. PMID 11301255. S2CID 14670796.
- ^ Rosario CO, Ko MA, Haffani YZ, Gladdy RA, Paderova J, Pollett A, Squire JA, Dennis JW, Swallow CJ (April 2010). "Plk4 is required for cytokinesis and maintenance of chromosomal stability". Proceedings of the National Academy of Sciences of the United States of America. 107 (15): 6888–93. Bibcode:2010PNAS..107.6888R. doi:10.1073/pnas.0910941107. PMC 2872425. PMID 20348415.
- ^ Puklowski A, Homsi Y, Keller D, May M, Chauhan S, Kossatz U, Grünwald V, Kubicka S, Pich A, Manns MP, Hoffmann I, Gönczy P, Malek NP (July 2011). "The SCF-FBXW5 E3-ubiquitin ligase is regulated by PLK4 and targets HsSAS-6 to control centrosome duplication". Nature Cell Biology. 13 (8): 1004–9. doi:10.1038/ncb2282. PMID 21725316. S2CID 24052533.
- ^ Cunha-Ferreira I, Bento I, Pimenta-Marques A, Jana SC, Lince-Faria M, Duarte P, Borrego-Pinto J, Gilberto S, Amado T, Brito D, Rodrigues-Martins A, Debski J, Dzhindzhev N, Bettencourt-Dias M (November 2013). "Regulation of autophosphorylation controls PLK4 self-destruction and centriole number". Current Biology. 23 (22): 2245–54. Bibcode:2013CBio...23.2245C. doi:10.1016/j.cub.2013.09.037. hdl:10400.7/833. PMID 24184099. S2CID 9056074.
- ^ Guderian G, Westendorf J, Uldschmid A, Nigg EA (July 2010). "Plk4 trans-autophosphorylation regulates centriole number by controlling betaTrCP-mediated degradation" (PDF). Journal of Cell Science. 123 (Pt 13): 2163–9. doi:10.1242/jcs.068502. PMID 20516151. S2CID 14815516.
- ^ Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, Klitgord N, Simon C, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Albala JS, Lim J, Fraughton C, Llamosas E, Cevik S, Bex C, Lamesch P, Sikorski RS, Vandenhaute J, Zoghbi HY, Smolyar A, Bosak S, Sequerra R, Doucette-Stamm L, Cusick ME, Hill DE, Roth FP, Vidal M (October 2005). "Towards a proteome-scale map of the human protein-protein interaction network". Nature. 437 (7062): 1173–8. Bibcode:2005Natur.437.1173R. doi:10.1038/nature04209. PMID 16189514. S2CID 4427026.
Further reading
- Kleylein-Sohn J, Westendorf J, Le Clech M, Habedanck R, Stierhof YD, Nigg EA (August 2007). "Plk4-induced centriole biogenesis in human cells" (PDF). Developmental Cell. 13 (2): 190–202. doi:10.1016/j.devcel.2007.07.002. PMID 17681131.
- Bettencourt-Dias M, Rodrigues-Martins A, Carpenter L, Riparbelli M, Lehmann L, Gatt MK, Carmo N, Balloux F, Callaini G, Glover DM (December 2005). "SAK/PLK4 is required for centriole duplication and flagella development" (PDF). Current Biology. 15 (24): 2199–207. Bibcode:2005CBio...15.2199B. doi:10.1016/j.cub.2005.11.042. PMID 16326102. S2CID 1257892.
- Habedanck R, Stierhof YD, Wilkinson CJ, Nigg EA (November 2005). "The Polo kinase Plk4 functions in centriole duplication". Nature Cell Biology. 7 (11): 1140–6. doi:10.1038/ncb1320. PMID 16244668. S2CID 1349505.
- Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, Klitgord N, Simon C, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Albala JS, Lim J, Fraughton C, Llamosas E, Cevik S, Bex C, Lamesch P, Sikorski RS, Vandenhaute J, Zoghbi HY, Smolyar A, Bosak S, Sequerra R, Doucette-Stamm L, Cusick ME, Hill DE, Roth FP, Vidal M (October 2005). "Towards a proteome-scale map of the human protein-protein interaction network". Nature. 437 (7062): 1173–8. Bibcode:2005Natur.437.1173R. doi:10.1038/nature04209. PMID 16189514. S2CID 4427026.
- Li J, Tan M, Li L, Pamarthy D, Lawrence TS, Sun Y (April 2005). "SAK, a new polo-like kinase, is transcriptionally repressed by p53 and induces apoptosis upon RNAi silencing". Neoplasia. 7 (4): 312–23. doi:10.1593/neo.04325. PMC 1501148. PMID 15967108.
- Barrios-Rodiles M, Brown KR, Ozdamar B, Bose R, Liu Z, Donovan RS, Shinjo F, Liu Y, Dembowy J, Taylor IW, Luga V, Przulj N, Robinson M, Suzuki H, Hayashizaki Y, Jurisica I, Wrana JL (March 2005). "High-throughput mapping of a dynamic signaling network in mammalian cells". Science. 307 (5715): 1621–5. Bibcode:2005Sci...307.1621B. doi:10.1126/science.1105776. PMID 15761153. S2CID 39457788.
- Suzuki Y, Yamashita R, Shirota M, Sakakibara Y, Chiba J, Mizushima-Sugano J, Nakai K, Sugano S (September 2004). "Sequence comparison of human and mouse genes reveals a homologous block structure in the promoter regions". Genome Research. 14 (9): 1711–8. doi:10.1101/gr.2435604. PMC 515316. PMID 15342556.
- Macmillan JC, Hudson JW, Bull S, Dennis JW, Swallow CJ (October 2001). "Comparative expression of the mitotic regulators SAK and PLK in colorectal cancer". Annals of Surgical Oncology. 8 (9): 729–40. doi:10.1007/s10434-001-0729-6. PMID 11597015. S2CID 21516819.
- Yamashita Y, Kajigaya S, Yoshida K, Ueno S, Ota J, Ohmine K, Ueda M, Miyazato A, Ohya K, Kitamura T, Ozawa K, Mano H (October 2001). "Sak serine-threonine kinase acts as an effector of Tec tyrosine kinase". The Journal of Biological Chemistry. 276 (42): 39012–20. doi:10.1074/jbc.M106249200. PMID 11489907.
- Hudson JW, Chen L, Fode C, Binkert C, Dennis JW (January 2000). "Sak kinase gene structure and transcriptional regulation". Gene. 241 (1): 65–73. doi:10.1016/S0378-1119(99)00467-9. PMID 10607900.
- Schultz SJ, Nigg EA (October 1993). "Identification of 21 novel human protein kinases, including 3 members of a family related to the cell cycle regulator nimA of Aspergillus nidulans". Cell Growth & Differentiation. 4 (10): 821–30. PMID 8274451.
- Man CH, Lam W, Dang CC, Zeng XY, Zheng LC, Chan NN, Ng KL, Chan KC, Kwok TH, Ng TC, Leung WY, Huen MS, Wong CC, So CW, Dou Z, Goyama S, Bray MR, Mak TW, Leung AY (December 2023). "Inhibition of PLK4 remodels histone methylation and activates the immune response via the cGAS-STING pathway in TP53-mutated AML". Blood. 142 (23): 2002–2015. doi:10.1182/blood.2023019782. PMID 37738460.
External links
- Overview of all the structural information available in the PDB for UniProt: O00444 (Human Serine/threonine-protein kinase PLK4) at the PDBe-KB.
- Overview of all the structural information available in the PDB for UniProt: Q64702 (Mouse Serine/threonine-protein kinase PLK4) at the PDBe-KB.