
Exome sequencing, also known as whole exome sequencing (WES), is a genomic technique for sequencing all of the protein-coding regions of genes in a genome (known as the exome).[1] It consists of two steps: the first step is to select only the subset of DNA that encodes proteins. These regions are known as exons—humans have about 180,000 exons, constituting about 1% of the human genome, or approximately 30 million base pairs. The second step is to sequence the exonic DNA using any high-throughput DNA sequencing technology.[2]
The goal of this approach is to identify genetic variants that alter protein sequences, and to do this at a much lower cost than whole-genome sequencing. Since these variants can be responsible for both Mendelian and common polygenic diseases, such as Alzheimer's disease, whole exome sequencing has been applied both in academic research and as a clinical diagnostic.

Motivation and comparison to other approaches
Exome sequencing is especially effective in the study of rare Mendelian diseases, because it is an efficient way to identify the genetic variants in all of an individual's genes. These diseases are most often caused by very rare genetic variants that are only present in a tiny number of individuals;[3] by contrast, techniques such as SNP arrays can only detect shared genetic variants that are common to many individuals in the wider population.[4] Furthermore, because severe disease-causing variants are much more likely (but by no means exclusively) to be in the protein coding sequence,[5][6] focusing on this 1% costs far less than whole genome sequencing but still detects a high yield of relevant variants.
In the past, clinical genetic tests were chosen based on the clinical presentation of the patient (i.e. focused on one gene or a small number known to be associated with a particular syndrome), or surveyed only certain types of variation (e.g. comparative genomic hybridization) but provided definitive genetic diagnoses in fewer than half of all patients.[7] Exome sequencing is now increasingly used to complement these other tests: both to find mutations in genes already known to cause disease as well as to identify novel genes by comparing exomes from patients with similar features.[citation needed]
Technical methodology
Step 1: Target-enrichment strategies
Target-enrichment methods allow one to selectively capture genomic regions of interest from a DNA sample prior to sequencing. Several target-enrichment strategies have been developed since the original description of the direct genomic selection (DGS) method in 2005.[8]
Though many techniques have been described for targeted capture, only a few of these have been extended to capture entire exomes.[9] The first target enrichment strategy to be applied to whole exome sequencing was the array-based hybrid capture method in 2007, but in-solution capture has gained popularity in recent years.
Array-based capture
Microarrays contain single-stranded oligonucleotides with sequences from the human genome to tile the region of interest fixed to the surface. Genomic DNA is sheared to form double-stranded fragments. The fragments undergo end-repair to produce blunt ends and adaptors with universal priming sequences are added. These fragments are hybridized to oligos on the microarray. Unhybridized fragments are washed away and the desired fragments are eluted. The fragments are then amplified using PCR.[10][11]
Roche NimbleGen was first to take the original DGS technology[8] and adapt it for next-generation sequencing. They developed the Sequence Capture Human Exome 2.1M Array to capture ~180,000 coding exons.[12] This method is both time-saving and cost-effective compared to PCR based methods. The Agilent Capture Array and the comparative genomic hybridization array are other methods that can be used for hybrid capture of target sequences. Limitations in this technique include the need for expensive hardware as well as a relatively large amount of DNA.[13]
In-solution capture
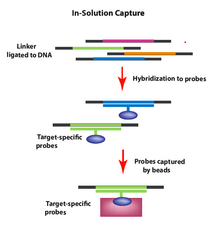
To capture genomic regions of interest using in-solution capture, a pool of custom oligonucleotides (probes) is synthesized and hybridized in solution to a fragmented genomic DNA sample. The probes (labeled with beads) selectively hybridize to the genomic regions of interest after which the beads (now including the DNA fragments of interest) can be pulled down and washed to clear excess material. The beads are then removed and the genomic fragments can be sequenced allowing for selective DNA sequencing of genomic regions (e.g., exons) of interest.
This method was developed to improve on the hybridization capture target-enrichment method. In solution capture (as opposed to hybrid capture) there is an excess of probes to target regions of interest over the amount of template required.[13] The optimal target size is about 3.5 megabases and yields excellent sequence coverage of the target regions. The preferred method is dependent on several factors including: number of base pairs in the region of interest, demands for reads on target, equipment in house, etc.[14]
Step 2: Sequencing
There are many Next Generation Sequencing sequencing platforms available, postdating classical Sanger sequencing methodologies. Other platforms include Roche 454 sequencer and Life Technologies SOLiD systems, the Life Technologies Ion Torrent and Illumina's Illumina Genome Analyzer II (defunct) and subsequent Illumina MiSeq, HiSeq, and NovaSeq series instruments, all of which can be used for massively parallel exome sequencing. These 'short read' NGS systems are particularly well suited to analyse many relatively short stretches of DNA sequence, as found in human exons.
Comparison with other technologies
There are multiple technologies available that identify genetic variants. Each technology has advantages and disadvantages in terms of technical and financial factors. Two such technologies are microarrays and whole-genome sequencing.
Microarray-based genotyping
Microarrays use hybridization probes to test the prevalence of known DNA sequences, thus they cannot be used to identify unexpected genetic changes.[13] In contrast, the high-throughput sequencing technologies used in exome sequencing directly provide the nucleotide sequences of DNA at the thousands of exonic loci tested.[15] Hence, WES addresses some of the present limitations of hybridization genotyping arrays.
Although exome sequencing is more expensive than hybridization-based technologies on a per-sample basis, its cost has been decreasing due to the falling cost and increased throughput of whole genome sequencing.[citation needed]
Whole-genome sequencing
Exome sequencing is only able to identify those variants found in the coding region of genes which affect protein function. It is not able to identify the structural and non-coding variants associated with the disease, which can be found using other methods such as whole genome sequencing.[2] There remains 99% of the human genome that is not covered using exome sequencing, and exome sequencing allows sequencing of portions of the genome over at least 20 times as many samples compared to whole genome sequencing.[2] For translation of identified rare variants into the clinic, sample size and the ability to interpret the results to provide a clinical diagnosis indicates that with the current knowledge in genetics, there are reports of exome sequencing being used for assisting diagnosis.[12] The cost of exome sequencing is typically lower than whole genome sequencing.[16]
Data analysis
The statistical analysis of the large quantity of data generated from sequencing approaches is a challenge. Even by only sequencing the exomes of individuals, a large quantity of data and sequence information is generated which requires a significant amount of data analysis. Challenges associated with the analysis of this data include changes in programs used to align and assemble sequence reads.[13] Various sequencing technologies also have different error rates and generate various read-lengths which can pose challenges in comparing results from different sequencing platforms.
False positive and false negative findings are associated with genomic resequencing approaches and are critical issues. A few strategies have been developed to improve the quality of exome data such as:
- Comparing the genetic variants identified between sequencing and array-based genotyping[2]
- Comparing the coding SNPs to a whole genome sequenced individual with the disorder[2]
- Comparing the coding SNPs with Sanger sequencing of HapMap individuals[2]
Rare recessive disorders may not have single nucleotide polymorphisms (SNPs) in public databases such as dbSNP. More common recessive phenotypes would be more likely to have disease-causing variants reported in dbSNP. For example, the most common cystic fibrosis variant has an allele frequency of about 3% in most populations. Screening out such variants might erroneously exclude such genes from consideration. Genes for recessive disorders are usually easier to identify than dominant disorders because the genes are less likely to have more than one rare nonsynonymous variant.[2] The system that screens common genetic variants relies on dbSNP which may not have accurate information about the variation of alleles. Using lists of common variation from a study exome or genome-wide sequenced individual would be more reliable. A challenge in this approach is that as the number of exomes sequenced increases, dbSNP will also increase in the number of uncommon variants. It will be necessary to develop thresholds to define the common variants that are unlikely to be associated with a disease phenotype.[15]
Genetic heterogeneity and population ethnicity are also major limitations as they may increase the number of false positive and false negative findings which will make the identification of candidate genes more difficult. Of course, it is possible to reduce the stringency of the thresholds in the presence of heterogeneity and ethnicity, however this will reduce the power to detect variants as well. Using a genotype-first approach to identify candidate genes might also offer a solution to overcome these limitations.
Unlike common variant analysis, the analysis of rare variants in whole-exome sequencing studies evaluates variant sets rather than single variants.[17][18] Functional annotations predict the effect or function of rare variants and help prioritize rare functional variants. Incorporating these annotations can effectively boost the power of genetic association of rare variants analysis of whole genome sequencing studies.[19] Some methods and tools have been developed to perform functionally-informed rare variant association analysis by incorporating functional annotations to empower analysis in whole exome sequencing studies.[20][21]
Ethical implications
New technologies in genomics have changed the way researchers approach both basic and translational research. With approaches such as exome sequencing, it is possible to significantly enhance the data generated from individual genomes which has put forth a series of questions on how to deal with the vast amount of information. Should the individuals in these studies be allowed to have access to their sequencing information? Should this information be shared with insurance companies? This data can lead to unexpected findings and complicate clinical utility and patient benefit. This area of genomics still remains a challenge and researchers are looking into how to address these questions.[15]
Applications of exome sequencing
By using exome sequencing, fixed-cost studies can sequence samples to much higher depth than could be achieved with whole genome sequencing. This additional depth makes exome sequencing well suited to several applications that need reliable variant calls.
Rare variant mapping in complex disorders
Current association studies have focused on common variation across the genome, as these are the easiest to identify with our current assays. However, disease-causing variants of large effect have been found to lie within exomes in candidate gene studies, and because of negative selection, are found in much lower allele frequencies and may remain untyped in current standard genotyping assays. Whole genome sequencing is a potential method to assay novel variant across the genome. However, in complex disorders (such as autism), a large number of genes are thought to be associated with disease risk.[1][22] This heterogeneity of underlying risk means that very large sample sizes are required for gene discovery, and thus whole genome sequencing is not particularly cost-effective. This sample size issue is alleviated by the development of novel advanced analytic methods, which effectively map disease genes despite the genetic mutations are rare at variant level.[22] In addition, variants in coding regions have been much more extensively studied and their functional implications are much easier to derive, making the practical applications of variants within the targeted exome region more immediately accessible.
Exome sequencing in rare variant gene discovery remains a very active and ongoing area of research, and there is growing evidence that a significant burden of risk is observed across sets of genes. The exome sequencing has been reported rare variants in KRT82 gene in the autoimmune disorder Alopecia Areata.[1]
Discovery of Mendelian disorders
In Mendelian disorders of large effect, findings thus far suggest one or a very small number of variants within coding genes underlie the entire condition. Because of the severity of these disorders, the few causal variants are presumed to be extremely rare or novel in the population, and would be missed by any standard genotyping assay. Exome sequencing provides high coverage variant calls across coding regions, which are needed to separate true variants from noise. A successful model of Mendelian gene discovery involves the discovery of de novo variants using trio sequencing, where parents and proband are genotyped.
Case studies
A study published in September 2009 discussed a proof of concept experiment to determine if it was possible to identify causal genetic variants using exome sequencing. They sequenced four individuals with Freeman–Sheldon syndrome (FSS) (OMIM 193700), a rare autosomal dominant disorder known to be caused by a mutation in the gene MYH3.[2] Eight HapMap individuals were also sequenced to remove common variants in order to identify the causal gene for FSS. After exclusion of common variants, the authors were able to identify MYH3, which confirms that exome sequencing can be used to identify causal variants of rare disorders.[2] This was the first reported study that used exome sequencing as an approach to identify an unknown causal gene for a rare mendelian disorder.
Subsequently, another group reported successful clinical diagnosis of a suspected Bartter syndrome patient of Turkish origin.[12] Bartter syndrome is a renal salt-wasting disease. Exome sequencing revealed an unexpected well-conserved recessive mutation in a gene called SLC26A3 which is associated with congenital chloride diarrhea (CLD). This molecular diagnosis of CLD was confirmed by the referring clinician. This example provided proof of concept of the use of whole-exome sequencing as a clinical tool in evaluation of patients with undiagnosed genetic illnesses. This report is regarded as the first application of next generation sequencing technology for molecular diagnosis of a patient.
A second report was conducted on exome sequencing of individuals with a mendelian disorder known as Miller syndrome (MIM#263750), a rare disorder of autosomal recessive inheritance. Two siblings and two unrelated individuals with Miller syndrome were studied. They looked at variants that have the potential to be pathogenic such as non-synonymous mutations, splice acceptor and donor sites and short coding insertions or deletions.[3] Since Miller syndrome is a rare disorder, it is expected that the causal variant has not been previously identified. Previous exome sequencing studies of common single nucleotide polymorphisms (SNPs) in public SNP databases were used to further exclude candidate genes. After exclusion of these genes, the authors found mutations in DHODH that were shared among individuals with Miller syndrome. Each individual with Miller syndrome was a compound heterozygote for the DHODH mutations which were inherited as each parent of an affected individual was found to be a carrier.[3]
This was the first time exome sequencing was shown to identify a novel gene responsible for a rare mendelian disease. This exciting finding demonstrates that exome sequencing has the potential to locate causative genes in complex diseases, which previously has not been possible due to limitations in traditional methods. Targeted capture and massively parallel sequencing represents a cost-effective, reproducible and robust strategy with high sensitivity and specificity to detect variants causing protein-coding changes in individual human genomes.
Clinical diagnostics
Exome sequencing can be used to diagnose the genetic cause of disease in a patient. Identification of the underlying disease gene mutation(s) can have major implications for diagnostic and therapeutic approaches, can guide prediction of disease natural history, and makes it possible to test at-risk family members.[2][3][12][23][24][25] There are many factors that make exome sequencing superior to single gene analysis including the ability to identify mutations in genes that were not tested due to an atypical clinical presentation[25] or the ability to identify clinical cases where mutations from different genes contribute to the different phenotypes in the same patient.[3]
Having diagnosed a genetic cause of a disease, this information may guide the selection of appropriate treatment. The first time this strategy was performed successfully in the clinic was in the treatment of an infant with inflammatory bowel disease.[24][26] A number of conventional diagnostics had previously been used, but the results could not explain the infant's symptoms. Analysis of exome sequencing data identified a mutation in the XIAP gene. Knowledge of this gene's function guided the infant's treatment, leading to a bone marrow transplantation which cured the child of disease.[24]
Researchers have used exome sequencing to identify the underlying mutation for a patient with Bartter Syndrome and congenital chloride diarrhea.[12] Bilgular's group also used exome sequencing and identified the underlying mutation for a patient with severe brain malformations, stating "[These findings] highlight the use of whole exome sequencing to identify disease loci in settings in which traditional methods have proved challenging... Our results demonstrate that this technology will be particularly valuable for gene discovery in those conditions in which mapping has been confounded by locus heterogeneity and uncertainty about the boundaries of diagnostic classification, pointing to a bright future for its broad application to medicine".[23]
Researchers at University of Cape Town, South Africa used exome sequencing to discover the genetic mutation of CDH2 as the underlying cause of a genetic disorder known as arrhythmogenic right ventricle cardiomyopathy (ARVC)‚ which increases the risk of heart disease and cardiac arrest. [1]
Commercial costs
Multiple companies have offered exome sequencing to consumers. Knome was the first company to offer exome sequencing services to consumers[when?], at a cost of several thousand dollars.[27] Later, 23andMe ran a pilot WES program that was announced in September 2011 and was discontinued in 2012. Consumers could obtain exome data at a cost of $999. The company provided raw data, and did not offer analysis.[27][28][29]
In November 2012, DNADTC, a division of Gene by Gene started offering exomes at 80X coverage and introductory price of $695.[30] This price per DNADTC web site is currently $895. In October 2013, BGI announced a promotion for personal whole exome sequencing at 50X coverage for $499.[31] In June 2016 Genos was able to achieve an even lower price of $399 with a CLIA-certified 75X consumer exome sequenced from saliva.[32][33][34]
A 2018 review of 36 studies found the cost for exome sequencing to range from $555 USD to $5,169 USD, with a diagnostic yield ranging from 3% to 79% depending on patient groups.[16]
See also
References
- ^ a b c Erjavec SO, Gelfman S, Abdelaziz AR, Lee EY, Monga I, Alkelai A, Ionita-Laza I, Petukhova L, Christiano AM (Feb 2022). "Whole exome sequencing in Alopecia Areata identifies rare variants in KRT82". Nat Commun. 13 (1): 800. Bibcode:2022NatCo..13..800E. doi:10.1038/s41467-022-28343-3. PMC 8831607. PMID 35145093.
- ^ a b c d e f g h i j Ng SB, Turner EH, Robertson PD, Flygare SD, Bigham AW, Lee C, Shaffer T, Wong M, Bhattacharjee A, Eichler EE, Bamshad M, Nickerson DA, Shendure J (10 September 2009). "Targeted capture and massively parallel sequencing of 12 human exomes". Nature. 461 (7261): 272–276. Bibcode:2009Natur.461..272N. doi:10.1038/nature08250. PMC 2844771. PMID 19684571.
- ^ a b c d e Sarah B Ng; Kati J Buckingham; Choli Lee; Abigail W Bigham; Holly K Tabor; Karin M Dent; Chad D Huff; Paul T Shannon; Ethylin Wang Jabs; Deborah A Nickerson; Jay Shendure; Michael J Bamshad (2010). "Exome sequencing identifies the cause of a mendelian disorder". Nature Genetics. 42 (1): 30–35. doi:10.1038/ng.499. PMC 2847889. PMID 19915526.
- ^ Wang, D. G.; Fan, J. B.; Siao, C. J.; Berno, A.; Young, P.; Sapolsky, R.; Ghandour, G.; Perkins, N.; Winchester, E. (1998-05-15). "Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome". Science. 280 (5366): 1077–1082. Bibcode:1998Sci...280.1077W. doi:10.1126/science.280.5366.1077. ISSN 0036-8075. PMID 9582121.
- ^ Petersen, Britt-Sabina; Fredrich, Broder; Hoeppner, Marc P.; Ellinghaus, David; Franke, Andre (2017-02-14). "Opportunities and challenges of whole-genome and -exome sequencing". BMC Genetics. 18 (1): 14. doi:10.1186/s12863-017-0479-5. ISSN 1471-2156. PMC 5307692. PMID 28193154.
- ^ Botstein, David; Risch, Neil (March 2003). "Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease". Nature Genetics. 33 (3): 228–237. doi:10.1038/ng1090. ISSN 1546-1718. PMID 12610532. S2CID 10599219.
- ^ Rauch, A; Hoyer, J; Guth, S; Zweier, C; Kraus, C; Becker, C; Zenker, M; Hüffmeier, U; Thiel, C; Rüschendorf, F; Nürnberg, P; Reis, A; Trautmann, U (Oct 1, 2006). "Diagnostic yield of various genetic approaches in patients with unexplained developmental delay or mental retardation". American Journal of Medical Genetics Part A. 140 (19): 2063–74. doi:10.1002/ajmg.a.31416. PMID 16917849. S2CID 24570999.
- ^ a b Stavros Basiardes; Rose Veile; Cindy Helms; Elaine R. Mardis; Anne M. Bowcock; Michael Lovett (2005). "Direct Genomic Selection". Nature Methods. 1 (2): 63–69. doi:10.1038/nmeth0105-63. PMID 16152676. S2CID 667227.
- ^ Teer, J. K.; Mullikin, J. C. (12 August 2010). "Exome sequencing: the sweet spot before whole genomes". Human Molecular Genetics. 19 (R2): R145–R151. doi:10.1093/hmg/ddq333. PMC 2953745. PMID 20705737.
- ^ Emily H. Turner; Sarah B. Ng; Deborah A. Nickerson; Jay Shendure (2009). "Methods for Genomic Partitioning". Annu Rev Genom Hum Genet. 10 (1): 30–35. doi:10.1146/annurev-genom-082908-150112. PMID 19630561.
- ^ Mertes F, Elsharawy A, Sauer S, van Helvoort JM, van der Zaag PJ, Franke A, Nilsson M, Lehrach H, Brookes AJ (2011). "Targeted enrichment of genomic DNA regions for next-generation sequencing". Brief. Funct. Genomics. 10 (6): 374–386. doi:10.1093/bfgp/elr033. PMC 3245553. PMID 22121152. Archived from the original (PDF) on 2017-09-21. Retrieved 2018-10-24.
- ^ a b c d e Choi M, Scholl UI, Ji W, Liu T, Tikhonova IR, Zumbo P, Nayir A, Bakkaloğlu A, Ozen S, Sanjad S, Nelson-Williams C, Farhi A, Mane S, Lifton RP (10 November 2009). "Genetic diagnosis by whole exome capture and massively parallel DNA sequencing". Proc Natl Acad Sci U S A. 106 (45): 19096–19101. Bibcode:2009PNAS..10619096C. doi:10.1073/pnas.0910672106. PMC 2768590. PMID 19861545.
- ^ a b c d Kahvejian A, Quackenbush J, Thompson JF (2008). "What would you do if you could sequence everything?". Nature Biotechnology. 26 (10): 1125–1133. doi:10.1038/nbt1494. PMC 4153598. PMID 18846086.
- ^ Lira Mamanova; Coffey, Alison J; Scott, Carol E; Kozarewa, Iwanka; Turner, Emily H; Kumar, Akash; Howard, Eleanor; Shendure, Jay; Turner, Daniel J; et al. (February 2010). "Target-enrichment strategies for nextgeneration sequencing". Nature Methods. 7 (2): 111–118. doi:10.1038/nmeth.1419. PMID 20111037. S2CID 21410733.
- ^ a b c Biesecker LG (Jan 2010). "Exome sequencing makes medical genomics a reality". Nat. Genet. 42 (1): 13–14. doi:10.1038/ng0110-13. PMID 20037612.
- ^ a b Schwarze, K; Buchanan, J; Taylor, JC; Wordsworth, S (May 2018). "Are whole Exome and whole Genome Sequencing Approaches Cost-Effective? A Systematic Review of the Literature". Value in Health. 21: S100. doi:10.1016/j.jval.2018.04.677. ISSN 1098-3015.
- ^ Lee, Seunggeung; Abecasis, Goncalo R.; Boehnke, Michael; Lin, Xihong (July 2014). "Rare-Variant Association Analysis: Study Designs and Statistical Tests". The American Journal of Human Genetics. 95 (1): 5–23. doi:10.1016/j.ajhg.2014.06.009. PMC 4085641. PMID 24995866.
- ^ Wang, Quanli; Dhindsa, Ryan S.; Carss, Keren; Harper, Andrew R.; et al. (23 September 2021). "Rare variant contribution to human disease in 281,104 UK Biobank exomes". Nature. 597 (7877): 527–532. Bibcode:2021Natur.597..527W. doi:10.1038/s41586-021-03855-y. PMC 8458098. PMID 34375979.
- ^ Li, Xihao; Li, Zilin; Zhou, Hufeng; Gaynor, Sheila M.; et al. (September 2020). "Dynamic incorporation of multiple in silico functional annotations empowers rare variant association analysis of large whole-genome sequencing studies at scale". Nature Genetics. 52 (9): 969–983. doi:10.1038/s41588-020-0676-4. PMC 7483769. PMID 32839606.
- ^ "STAARpipeline: an all-in-one rare-variant tool for biobank-scale whole-genome sequencing data". Nature Methods. 19 (12): 1532–1533. December 2022. doi:10.1038/s41592-022-01641-w. PMID 36316564. S2CID 253246835.
- ^ Li, Xihao; Quick, Corbin; Zhou, Hufeng; Gaynor, Sheila M.; Liu, Yaowu; Chen, Han; Selvaraj, Margaret Sunitha; Sun, Ryan; Dey, Rounak; Arnett, Donna K.; Bielak, Lawrence F.; Bis, Joshua C.; Blangero, John; Boerwinkle, Eric; Bowden, Donald W.; Brody, Jennifer A.; Cade, Brian E.; Correa, Adolfo; Cupples, L. Adrienne; Curran, Joanne E.; de Vries, Paul S.; Duggirala, Ravindranath; Freedman, Barry I.; Göring, Harald H. H.; Guo, Xiuqing; Haessler, Jeffrey; Kalyani, Rita R.; Kooperberg, Charles; Kral, Brian G.; Lange, Leslie A.; Manichaikul, Ani; Martin, Lisa W.; McGarvey, Stephen T.; Mitchell, Braxton D.; Montasser, May E.; Morrison, Alanna C.; Naseri, Take; O'Connell, Jeffrey R.; Palmer, Nicholette D.; Peyser, Patricia A.; Psaty, Bruce M.; Raffield, Laura M.; Redline, Susan; Reiner, Alexander P.; Reupena, Muagututi'a Sefuiva; Rice, Kenneth M.; Rich, Stephen S.; Sitlani, Colleen M.; Smith, Jennifer A.; Taylor, Kent D.; Vasan, Ramachandran S.; Willer, Cristen J.; Wilson, James G.; Yanek, Lisa R.; Zhao, Wei; NHLBI Trans-Omics for Precision Medicine (TOPMed) Consortium; TOPMed Lipids Working Group; Rotter, Jerome I.; Natarajan, Pradeep; Peloso, Gina M.; Li, Zilin; Lin, Xihong (January 2023). "Powerful, scalable and resource-efficient meta-analysis of rare variant associations in large whole genome sequencing studies". Nature Genetics. 55 (1): 154–164. doi:10.1038/s41588-022-01225-6. PMC 10084891. PMID 36564505. S2CID 255084231.
- ^ a b Weijun Luo; Chaolin Zhang; Yong-hui Jiang; Cory R. Brouwer (2018). "Systematic reconstruction of autism biology from massive genetic mutation profiles". Science Advances. 4 (4): e1701799. Bibcode:2018SciA....4.1799L. doi:10.1126/sciadv.1701799. PMC 5895441. PMID 29651456.
- ^ a b Bilgüvar K, Oztürk AK, Louvi A, Kwan KY, Choi M, Tatli B, Yalnizoğlu D, Tüysüz B, Cağlayan AO, Gökben S, Kaymakçalan H, Barak T, Bakircioğlu M, Yasuno K, Ho W, Sanders S, Zhu Y, Yilmaz S, Dinçer A, Johnson MH, Bronen RA, Koçer N, Per H, Mane S, Pamir MN, Yalçinkaya C, Kumandaş S, Topçu M, Ozmen M, Sestan N, Lifton RP, State MW, Günel M (9 September 2010). "Whole-exome sequencing identifies recessive WDR62 mutations in severe brain malformations". Nature. 467 (7312): 207–210. Bibcode:2010Natur.467..207B. doi:10.1038/nature09327. PMC 3129007. PMID 20729831.
- ^ a b c Worthey EA, Mayer AN, Syverson GD, Helbling D, Bonacci BB, Decker B, Serpe JM, Dasu T, Tschannen MR, Veith RL, Basehore MJ, Broeckel U, Tomita-Mitchell A, Arca MJ, Casper JT, Margolis DA, Bick DP, Hessner MJ, Routes JM, Verbsky JW, Jacob HJ, Dimmock DP (Mar 2011). "Making a definitive diagnosis: successful clinical application of whole exome sequencing in a child with intractable inflammatory bowel disease". Genet. Med. 13 (3): 255–262. doi:10.1097/GIM.0b013e3182088158. PMID 21173700.
- ^ a b Raffan E, Hurst LA, Turki SA, Carpenter G, Scott C, Daly A, Coffey A, Bhaskar S, Howard E, Khan N, Kingston H, Palotie A, Savage DB, O'Driscoll M, Smith C, O'Rahilly S, Barroso I, Semple RK (2011). "Early Diagnosis of Werner's Syndrome Using Exome-Wide Sequencing in a Single, Atypical Patient". Front Endocrinol (Lausanne). 2 (8): 8. doi:10.3389/fendo.2011.00008. PMC 3356119. PMID 22654791.
- ^ Warr, A.; Robert, C.; Hume, D.; Archibald, A.; Deeb, N.; Watson, M. (2 July 2015). "Exome Sequencing: Current and Future Perspectives". G3. 5 (8): 1543–1550. doi:10.1534/g3.115.018564. PMC 4528311. PMID 26139844.
- ^ a b Herper, Matthew (27 September 2011). "The Future Is Now: 23andMe Now Offers All Your Genes For $999". Forbes. Retrieved 11 December 2011.
- ^ "23andMe Launches Pilot Program for Direct-to-Consumer Exome Sequencing". GenomeWeb. GenomeWeb LLC. 28 September 2011. Retrieved 11 December 2011.
- ^ "Personal Genome Sequencing in Ostensibly Healthy Individuals and the PeopleSeq Consortium" (PDF). Genomes2People. 14 June 2016.
- ^ Vorhaus, Dan (29 November 2012). "DNA DTC: The Return of Direct to Consumer Whole Genome Sequencing". Genomics Law Report. Retrieved 30 May 2013.
- ^ "Ultimate Exome Promotion". BGI Americas. 18 October 2013. Archived from the original on 10 November 2013. Retrieved 17 November 2013.
- ^ "Owning Your Data: The Genos Model". Bio-IT World. 6 July 2016.
- ^ "Consumer Genomics Startup Genos Research Plans to Let Customers Explore, Share Their Data". GenomeWeb LLC. 13 June 2016.
- ^ "Genos - Own your DNA, Learn about Yourself, Drive Research". www.genosresearch.com. Archived from the original on 2016-08-25. Retrieved 2016-10-18.








