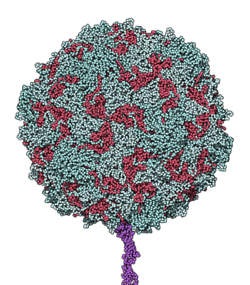
CD155 (cluster of differentiation 155), also known as the poliovirus receptor, is a protein that in humans is encoded by the PVR gene.[3][4] It is a transmembrane protein that is involved in forming junctions between neighboring cells. It is also the molecule that poliovirus uses to enter cells. The gene is specific to the primates.
Function
CD155 is a Type I transmembrane glycoprotein in the immunoglobulin superfamily.[5] Its normal cellular function is in the establishment of intercellular adherens junctions between epithelial cells.[6]
The external domain mediates cell attachment to the extracellular matrix molecule vitronectin, while its intracellular domain interacts with the dynein light chain Tctex-1/DYNLT1.
The role of CD155 in the immune system is unclear, though it may be involved in intestinal humoral immune responses.[6] Subsequent data has also suggested that CD155 may also be used to positively select MHC-independent T cells in the thymus.[citation needed]
Polio
Commonly known as Poliovirus Receptor (PVR), the protein serves as a cellular receptor for poliovirus in the first step of poliovirus replication. Transgenic mice that express the PVR gene have been constructed in order to study polio experimentally.[7]
Structure
CD155 is a transmembrane protein with 3 extracellular immunoglobulin-like domains, D1-D3, where D1 is recognized by the virus.[8]
Low resolution structures of CD155 complexed with poliovirus have been obtained using electron microscopy[9] while a high resolution structures of the ectodomain D1 and D2 of CD155 were solved by x-ray crystallography.[8]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000073008 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: poliovirus receptor".
- ^ Koike S, Horie H, Ise I, Okitsu A, Yoshida M, Iizuka N, Takeuchi K, Takegami T, Nomoto A (October 1990). "The poliovirus receptor protein is produced both as membrane-bound and secreted forms". EMBO J. 9 (10): 3217–24. doi:10.1002/j.1460-2075.1990.tb07520.x. PMC 552052. PMID 2170108.
- ^ Mendelsohn CL, Wimmer E, Racaniello VR (1989). "Cellular receptor for poliovirus: molecular cloning, nucleotide sequence, and expression of a new member of the immunoglobulin superfamily". Cell. 56 (5): 855–65. doi:10.1016/0092-8674(89)90690-9. PMID 2538245. S2CID 44296539.
- ^ a b Maier MK, Seth S, Czeloth N, et al. (2007). "The adhesion receptor CD155 determines the magnitude of humoral immune responses against orally ingested antigens". European Journal of Immunology. 37 (8): 2214–25. doi:10.1002/eji.200737072. PMID 17621371. S2CID 24583092.
- ^ Racaniello, Vincent R. (2006-01-05). "One hundred years of poliovirus pathogenesis". Virology. Virology 50th Anniversary Issue. 344 (1): 9–16. doi:10.1016/j.virol.2005.09.015. ISSN 0042-6822. PMID 16364730.
- ^ a b PDB: 3epc, 3epd, 3epf, 3eow; Zhang P, Mueller S, Morais MC, Bator CM, Bowman VD, Hafenstein S, Wimmer E, Rossmann MG (November 2008). "Crystal structure of CD155 and electron microscopic studies of its complexes with polioviruses". Proc. Natl. Acad. Sci. U.S.A. 105 (47): 18284–9. Bibcode:2008PNAS..10518284Z. doi:10.1073/pnas.0807848105. PMC 2587566. PMID 19011098.
- ^ PDB: 1DGI; He Y, Bowman VD, Mueller S, Bator CM, Bella J, Peng X, Baker TS, Wimmer E, Kuhn RJ, Rossmann MG (January 2000). "Interaction of the poliovirus receptor with poliovirus". Proc. Natl. Acad. Sci. U.S.A. 97 (1): 79–84. Bibcode:2000PNAS...97...79H. doi:10.1073/pnas.97.1.79. PMC 26619. PMID 10618374.
External links
- Human PVR genome location and PVR gene details page in the UCSC Genome Browser.
Further reading
- Pende D, Spaggiari GM, Marcenaro S, et al. (2005). "Analysis of the receptor-ligand interactions in the natural killer-mediated lysis of freshly isolated myeloid or lymphoblastic leukemias: evidence for the involvement of the Poliovirus receptor (CD155) and Nectin-2 (CD112)". Blood. 105 (5): 2066–73. doi:10.1182/blood-2004-09-3548. PMID 15536144.
- Pezzetti F, Palmieri A, Martinelli M, et al. (2007). "Linkage disequilibrium analysis of two genes mapping on OFC3: PVR and PVRL2". Eur. J. Hum. Genet. 15 (9): 992–4. doi:10.1038/sj.ejhg.5201868. PMID 17534374.
- Stanietsky N, Simic H, Arapovic J, et al. (2009). "The interaction of TIGIT with PVR and PVRL2 inhibits human NK cell cytotoxicity". Proc. Natl. Acad. Sci. U.S.A. 106 (42): 17858–63. Bibcode:2009PNAS..10617858S. doi:10.1073/pnas.0903474106. PMC 2764881. PMID 19815499.
- Tomasec P, Wang EC, Davison AJ, et al. (2005). "Downregulation of natural killer cell–activating ligand CD155 by human cytomegalovirus UL141". Nat. Immunol. 6 (2): 181–8. doi:10.1038/ni1156. PMC 2844263. PMID 15640804.
- Liu T, Qian WJ, Gritsenko MA, et al. (2005). "Human Plasma N-Glycoproteome Analysis by Immunoaffinity Subtraction, Hydrazide Chemistry, and Mass Spectrometry". J. Proteome Res. 4 (6): 2070–80. doi:10.1021/pr0502065. PMC 1850943. PMID 16335952.
- Kimura K, Wakamatsu A, Suzuki Y, et al. (2006). "Diversification of transcriptional modulation: Large-scale identification and characterization of putative alternative promoters of human genes". Genome Res. 16 (1): 55–65. doi:10.1101/gr.4039406. PMC 1356129. PMID 16344560.
- Pende D, Bottino C, Castriconi R, et al. (2005). "PVR (CD155) and Nectin-2 (CD112) as ligands of the human DNAM-1 (CD226) activating receptor: involvement in tumor cell lysis". Mol. Immunol. 42 (4): 463–9. doi:10.1016/j.molimm.2004.07.028. PMID 15607800.
- Kono T, Imai Y, Yasuda S, et al. (2008). "The CD155/poliovirus receptor enhances the proliferation of ras-mutated cells". Int. J. Cancer. 122 (2): 317–24. doi:10.1002/ijc.23080. PMID 17893876. S2CID 42542414.
- Minami Y, Ikeda W, Kajita M, et al. (2007). "Necl-5/poliovirus receptor interacts in cis with integrin alphaVbeta3 and regulates its clustering and focal complex formation". J. Biol. Chem. 282 (25): 18481–96. doi:10.1074/jbc.M611330200. PMID 17446174.
- Yu X, Harden K, Gonzalez LC, et al. (2009). "The surface protein TIGIT suppresses T cell activation by promoting the generation of mature immunoregulatory dendritic cells". Nat. Immunol. 10 (1): 48–57. doi:10.1038/ni.1674. PMID 19011627. S2CID 205361984.
- Ohka S, Igarashi H, Nagata N, et al. (2007). "Establishment of a Poliovirus Oral Infection System in Human Poliovirus Receptor-Expressing Transgenic Mice That Are Deficient in Alpha/Beta Interferon Receptor". J. Virol. 81 (15): 7902–12. doi:10.1128/JVI.02675-06. PMC 1951287. PMID 17507470.
- Zhang P, Mueller S, Morais MC, et al. (2008). "Crystal structure of CD155 and electron microscopic studies of its complexes with polioviruses". Proc. Natl. Acad. Sci. U.S.A. 105 (47): 18284–9. Bibcode:2008PNAS..10518284Z. doi:10.1073/pnas.0807848105. PMC 2587566. PMID 19011098.
- Gerhard DS, Wagner L, Feingold EA, et al. (2004). "The Status, Quality, and Expansion of the NIH Full-Length cDNA Project: The Mammalian Gene Collection (MGC)". Genome Res. 14 (10B): 2121–7. doi:10.1101/gr.2596504. PMC 528928. PMID 15489334.
- Carlsten M, Norell H, Bryceson YT, et al. (2009). "Primary human tumor cells expressing CD155 impair tumor targeting by down-regulating DNAM-1 on NK cells". J. Immunol. 183 (8): 4921–30. doi:10.4049/jimmunol.0901226. PMID 19801517.
- Kindberg E, Ax C, Fiore L, Svensson L (2009). "Ala67Thr mutation in the poliovirus receptor CD155 is a potential risk factor for vaccine and wild-type paralytic poliomyelitis". J. Med. Virol. 81 (5): 933–6. doi:10.1002/jmv.21444. PMID 19319949. S2CID 11567056.
- Bachelet I, Munitz A, Mankutad D, Levi-Schaffer F (2006). "Mast cell costimulation by CD226/CD112 (DNAM-1/Nectin-2): a novel interface in the allergic process". J. Biol. Chem. 281 (37): 27190–6. doi:10.1074/jbc.M602359200. PMID 16831868.
- Meyer D, Seth S, Albrecht J, et al. (2009). "CD96 interaction with CD155 via its first Ig-like domain is modulated by alternative splicing or mutations in distal Ig-like domains". J. Biol. Chem. 284 (4): 2235–44. doi:10.1074/jbc.M807698200. PMID 19056733.
- Pende D, Castriconi R, Romagnani P, et al. (2006). "Expression of the DNAM-1 ligands, Nectin-2 (CD112) and poliovirus receptor (CD155), on dendritic cells: relevance for natural killer-dendritic cell interaction". Blood. 107 (5): 2030–6. doi:10.1182/blood-2005-07-2696. hdl:2158/335995. PMID 16304049.
- Fujito T, Ikeda W, Kakunaga S, et al. (2005). "Inhibition of cell movement and proliferation by cell–cell contact-induced interaction of Necl-5 with nectin-3". J. Cell Biol. 171 (1): 165–73. doi:10.1083/jcb.200501090. PMC 2171219. PMID 16216929.
This article incorporates text from the United States National Library of Medicine, which is in the public domain.