| NCAM1 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | NCAM1, CD56, MSK39, NCAM, neural cell adhesion molecule 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 116930; MGI: 97281; HomoloGene: 40754; GeneCards: NCAM1; OMA:NCAM1 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Neural cell adhesion molecule (NCAM), also called CD56, is a homophilic binding glycoprotein expressed on the surface of neurons, glia and skeletal muscle. Although CD56 is often considered a marker of neural lineage commitment due to its discovery site, CD56 expression is also found in, among others, the hematopoietic system. Here, the expression of CD56 is mostly associated with, but not limited to, natural killer cells. CD56 has been detected on other lymphoid cells, including gamma delta (γδ) Τ cells and activated CD8+ T cells, as well as on dendritic cells.[5] NCAM has been implicated as having a role in cell–cell adhesion,[6] neurite outgrowth, synaptic plasticity, and learning and memory.
Forms, domains and homophilic binding
NCAM is a glycoprotein of Immunoglobulin (Ig) superfamily. At least 27 alternatively spliced NCAM mRNAs are produced, giving a wide diversity of NCAM isoforms.[7] The three main isoforms of NCAM vary only in their cytoplasmic domain:
- NCAM-120kDa (GPI anchored)
- NCAM-140kDa (short cytoplasmic domain)
- NCAM-180kDa (long cytoplasmic domain)
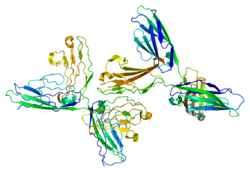
The extracellular domain of NCAM consists of five immunoglobulin-like (Ig) domains followed by two fibronectin type III (FNIII) domains. The different domains of NCAM have been shown to have different roles, with the Ig domains being involved in homophilic binding to NCAM, and the FNIII domains being involved signalling leading to neurite outgrowth.
Homophilic binding occurs between NCAM molecules on opposing surfaces (trans-) and NCAM molecules on the same surface (cis-)1. There is much controversy as to how exactly NCAM homophilic binding is arranged both in trans- and cis-. Current models suggest trans- homophilic binding occurs between two NCAM molecules binding antiparallel between all five Ig domains or just IgI and IgII. cis- homophilic binding is thought to occur by interactions between both IgI and IgII, and IgI and IgIII, forming a higher order NCAM multimer. Both cis- and trans- NCAM homophilic binding have been shown to be important in NCAM “activation” leading to neurite outgrowth.
Minor exons
Another layer of complexity is created by the insertion of other "minor" exons in the NCAM transcript. The two most notable are:
- the VASE (VAriable domain Spliced Exon) exon which is thought to correlate with an inhibition of the neurite outgrowth promoting properties of NCAM.
- the MSD (Muscle Specific Domain), which is thought to play a positive role in myoblast fusion.[8] In skeletal muscle it is found in all three NCAM isoforms, increasing their MW, giving NCAM-125, NCAM-145, and NCAM-185 isoforms, but is most commonly found in the NCAM-125 isoform.[8]
Posttranslational modification
NCAM exhibits glycoforms as it can be posttranslationally modified by the addition of polysialic acid (PSA) to the fifth Ig domain, which is thought to abrogate its homophilic binding properties and can lead to reduced cell adhesion important in cell migration and invasion. PSA has been shown to be critical in learning and memory. Removal of PSA from NCAM by the enzyme endoneuraminidase (EndoN) has been shown to abolish long-term potentiation (LTP) and long-term depression (LTD).[9][10][11]
Expression in normal cells
The neural cell adhesion molecule NCAM1 appears on early embryonic cells and is important in the formation of cell collectives and their boundaries at sites of morphogenesis.
Later in development, NCAM1 (CD56) expression is found on various differentiated tissues and is a major CAM mediating adhesion among neurons and between neurons and muscle.
Function
NCAM is thought to signal to induce neurite outgrowth via the fibroblast growth factor receptor (FGFR) and act upon the p59Fyn signaling pathway.
In nerves, NCAM1 regulates homophilic (like-like) interactions between neurons and between neurons and muscle; it associates with fibroblast growth factor receptor (FGFR) and stimulates tyrosine kinase activity of receptor to induce neurite outgrowth. When neural crest cells stop making N-CAM and N-cadherin, and start displaying integrin receptors, cells separate and migrate.
During hematopoiesis, CD56 is the prototypic marker of NK cells, also present on subset of CD4+ T cells and CD8+ T cells.
In cell adhesion, CD56 contributes to cell-cell adhesion or cell-matrix adhesion during embryonic development.
Pathology
In anatomic pathology, pathologists make use of CD56 immunohistochemistry to recognize certain tumors.
- Normal cells that stain positively for CD56 include NK cells, activated T cells, the brain and cerebellum, and neuroendocrine tissues.
- Tumors that are CD56-positive are myeloma, myeloid leukemia, neuroendocrine tumors, Wilms' tumor, neuroblastoma, extranodal NK/T-cell lymphoma, nasal type, pancreatic acinar cell carcinoma, pheochromocytoma, paraganglioma, small cell lung carcinoma, and the Ewing's sarcoma family of tumors.
Cancer
A member of the NCAM superfamily, NCAM2 gene has been observed progressively downregulated in human papillomavirus-positive neoplastic keratinocytes derived from uterine cervical preneoplastic lesions at different levels of malignancy.[12] For this reason, NCAM2 is likely to be associated with tumorigenesis and may be a potential prognostic marker for uterine cervical preneoplastic lesions progression.[12]
Alzheimer's disease
NCAM2 is found in lower levels in hippocampal synapses of Alzheimer's disease sufferers and is found to be broken down by beta-amyloid.[13]
Rabies
NCAM has been identified as one of the target proteins for the rabies virus, allowing entry into the cell.[14]
Anti-NCAM therapy
NCAM has been used as a target molecule for experimental antibody-based immunotherapy. Successful radio-immunolocalisation of metastases was demonstrated after giving injections of NCAM-binding 123J-UJ13a or 131J-UJ13a radio-immunoconjugates to children with neuroblastoma. Patients with small cell lung cancer were treated with the anti-NCAM immunotoxin huN901-DM1 in two different clinical studies, revealing acceptable toxicity and signs of clinical response.[15]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000149294 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000039542 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Van Acker HH, Capsomidis A, Smits EL, Van Tendeloo VF (2017). "CD56 in the Immune System: More Than a Marker for Cytotoxicity?". Frontiers in Immunology. 8: 892. doi:10.3389/fimmu.2017.00892. PMC 5522883. PMID 28791027.
- ^ Pathology Outlines
- ^ Reyes AA, Small SJ, Akeson R (March 1991). "At least 27 alternatively spliced forms of the neural cell adhesion molecule mRNA are expressed during rat heart development". Molecular and Cellular Biology. 11 (3): 1654–61. doi:10.1128/mcb.11.3.1654. PMC 369464. PMID 1996115.
- ^ a b Suzuki M, Angata K, Nakayama J, Fukuda M (December 2003). "Polysialic acid and mucin type o-glycans on the neural cell adhesion molecule differentially regulate myoblast fusion". The Journal of Biological Chemistry. 278 (49): 49459–68. doi:10.1074/jbc.M308316200. PMID 13679364.
- ^ Becker CG, Artola A, Gerardy-Schahn R, Becker T, Welzl H, Schachner M (July 1996). "The polysialic acid modification of the neural cell adhesion molecule is involved in spatial learning and hippocampal long-term potentiation". Journal of Neuroscience Research. 45 (2): 143–52. doi:10.1002/(SICI)1097-4547(19960715)45:2<143::AID-JNR6>3.0.CO;2-A. PMID 8843031. S2CID 43042018.
- ^ Stoenica L, Senkov O, Gerardy-Schahn R, Weinhold B, Schachner M, Dityatev A (May 2006). "In vivo synaptic plasticity in the dentate gyrus of mice deficient in the neural cell adhesion molecule NCAM or its polysialic acid". The European Journal of Neuroscience. 23 (9): 2255–64. doi:10.1111/j.1460-9568.2006.04771.x. PMID 16706834. S2CID 22798537.
- ^ Senkov O, Sun M, Weinhold B, Gerardy-Schahn R, Schachner M, Dityatev A (October 2006). "Polysialylated neural cell adhesion molecule is involved in induction of long-term potentiation and memory acquisition and consolidation in a fear-conditioning paradigm". The Journal of Neuroscience. 26 (42): 10888–109898. doi:10.1523/JNEUROSCI.0878-06.2006. PMC 6674738. PMID 17050727.
- ^ a b Rotondo JC, Bosi S, Bassi C, Ferracin M, Lanza G, Gafà R, Magri E, Selvatici R, Torresani S, Marci R, Garutti P, Negrini M, Tognon M, Martini F (April 2015). "Gene expression changes in progression of cervical neoplasia revealed by microarray analysis of cervical neoplastic keratinocytes". J Cell Physiol. 230 (4): 802–812. doi:10.1002/jcp.24808. hdl:11392/2066612. PMID 25205602. S2CID 24986454.
- ^ Leshchyns'ka I, Liew HT, Shepherd C, Halliday GM, Stevens CH, Ke YD, Ittner LM, Sytnyk V (November 2015). "Aβ-dependent reduction of NCAM2-mediated synaptic adhesion contributes to synapse loss in Alzheimer's disease". Nature Communications. 6 8836. Bibcode:2015NatCo...6.8836L. doi:10.1038/ncomms9836. PMC 4674770. PMID 26611261.
- ^ Guo Y, Duan M, Wang X, Gao J, Guan Z, Zhang M (April 2019). "Early events in rabies virus infection-Attachment, entry, and intracellular trafficking". Virus Research. 263: 217–225. doi:10.1016/j.virusres.2019.02.006. PMID 30772332. S2CID 73468336.
- ^ Jensen M, Berthold F (December 2007). "Targeting the neural cell adhesion molecule in cancer". Cancer Letters. 258 (1): 9–21. doi:10.1016/j.canlet.2007.09.004. PMID 17949897.
External links
- Neural+Cell+Adhesion+Molecule at the U.S. National Library of Medicine Medical Subject Headings (MeSH)













